Our Programs
LSD1
Lysine specific histone demethylase 1 (LSD1, also known as KDM1A) is an epigenetic “eraser”.
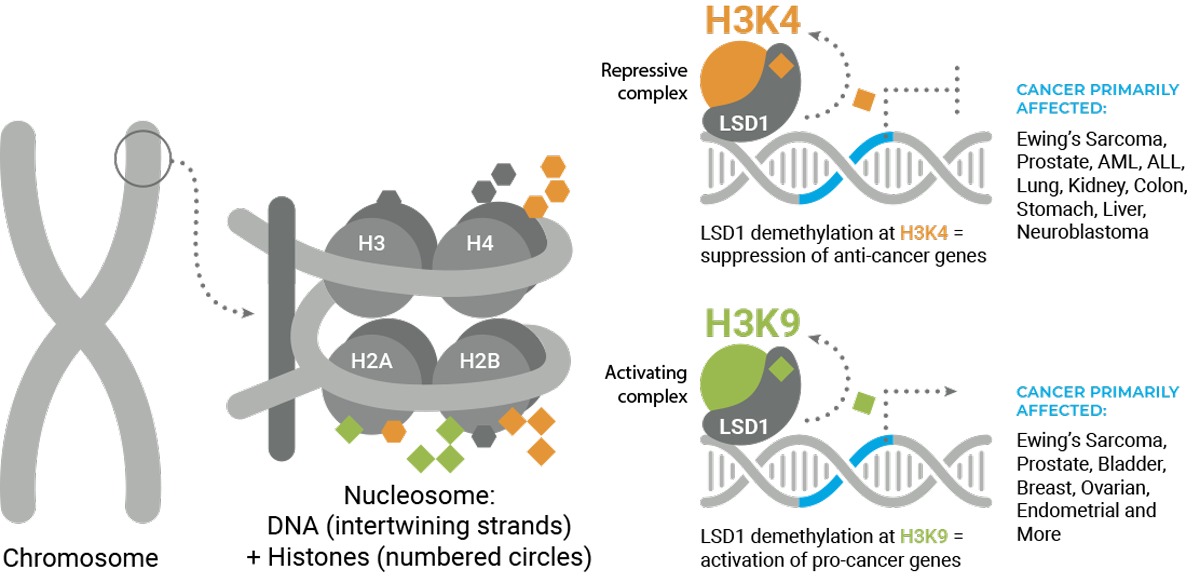
LSD1’s demethylation activity removes chemical groups, known as methyl marks, from protein-DNA structures known as nucleosomes. Nucleosomes comprise of DNA wrapped around structural proteins called histones. Histones, like most proteins, contain an amino acid known as lysine, a particularly important amino acid for chemical modification. LSD1 specifically erases methyl marks at lysine 4 and lysine 9 (H3K4, H3K9, “K” is the symbol for lysine). Depending on the presence or absence of methyl marks, DNA assumes an “open” conformation available for transcription of genes, or DNA assumes a “closed” conformation blocking gene expression.
A methyl mark at H3K4 is associated with an open conformation of DNA, and LSD1 closes DNA by removing the methyl mark. In contrast, a methyl mark at H3K9 is associated with a closed conformation of DNA, and LSD1 opens DNA by removing the methyl mark. LSD1 can therefore both promote and block gene expression, and is unique as an epigenetic modifier in this respect. Finally, each lysine can have up to three methyl marks, and LSD1 is only capable of removing the first two marks. The number of methyl marks can also have an effect on gene expression.
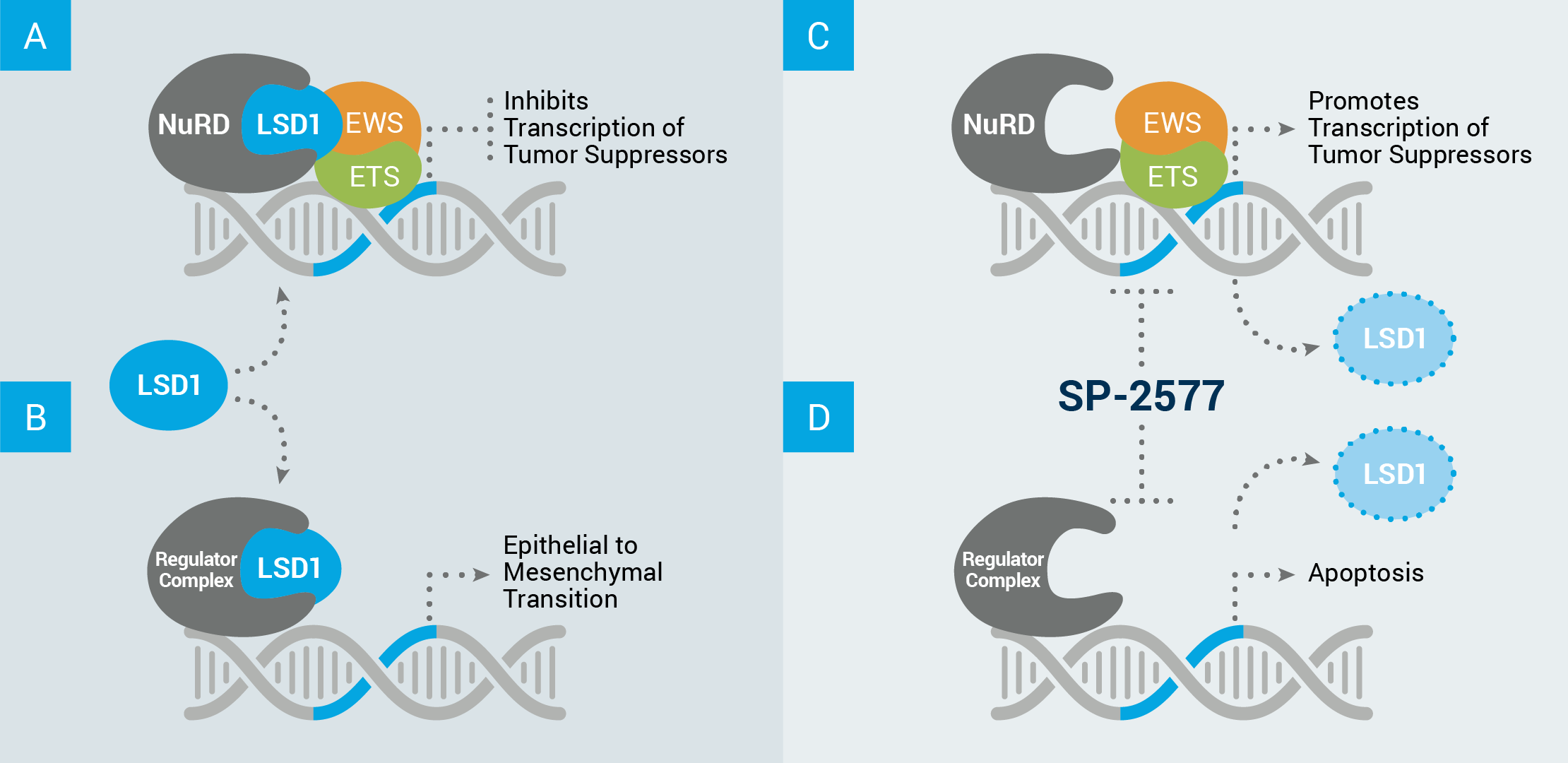
Above is an example of how LSD1’s scaffolding properties can drive cancer growth in Ewing sarcoma. (A) LSD1 acts in complex with other proteins, such as the nucleosome remodeling deacetylase complex (NuRD), to inhibit the transcription of tumor suppressor genes. (B) LSD1 can also interact with other proteins (ex: regulator complex) to disrupt the equilibrium of healthy cells and drive expression of a stem cell-like phenotype. This means cells that are differentiated, or designed to perform a tissue-specific function, revert to a more plastic, undefined state characterized by constant growth and invasion of nearby tissues. Seclidemstat inhibits LSD1’s scaffolding properties which (C) allows for the transcription of tumor suppressor genes, and/or (D) prevents transcription relating to self-renewal, and can cause cell apoptosis, i.e.,death.
In other cancers, LSD1 can associate with various other transcriptional co-activators and co-repressors. For instance, in prostate cancers LSD1 can interact with the androgen receptor to activate genes that drive tumor growth. For this reason, inhibiting LSD1’s demethylation and scaffolding properties is attractive for anti-tumor purposes across various cancer types.